1ysa: Difference between revisions
m Protected "1ysa" [edit=sysop:move=sysop] |
No edit summary |
||
| Line 1: | Line 1: | ||
[[Image: | ==THE GCN4 BASIC REGION LEUCINE ZIPPER BINDS DNA AS A DIMER OF UNINTERRUPTED ALPHA HELICES: CRYSTAL STRUCTURE OF THE PROTEIN-DNA COMPLEX== | ||
<StructureSection load='1ysa' size='340' side='right' caption='[[1ysa]], [[Resolution|resolution]] 2.90Å' scene=''> | |||
== Structural highlights == | |||
<table><tr><td colspan='2'>[[1ysa]] is a 4 chain structure with sequence from [http://en.wikipedia.org/wiki/Saccharomyces_cerevisiae Saccharomyces cerevisiae]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1YSA OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1YSA FirstGlance]. <br> | |||
</td></tr><tr><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1ysa FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1ysa OCA], [http://www.rcsb.org/pdb/explore.do?structureId=1ysa RCSB], [http://www.ebi.ac.uk/pdbsum/1ysa PDBsum]</span></td></tr> | |||
<table> | |||
== Evolutionary Conservation == | |||
[[Image:Consurf_key_small.gif|200px|right]] | |||
Check<jmol> | |||
<jmolCheckbox> | |||
<scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/ys/1ysa_consurf.spt"</scriptWhenChecked> | |||
<scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | |||
<text>to colour the structure by Evolutionary Conservation</text> | |||
</jmolCheckbox> | |||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/chain_selection.php?pdb_ID=2ata ConSurf]. | |||
<div style="clear:both"></div> | |||
<div style="background-color:#fffaf0;"> | |||
== Publication Abstract from PubMed == | |||
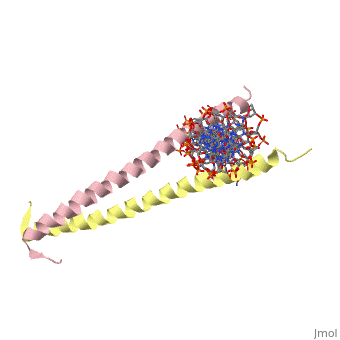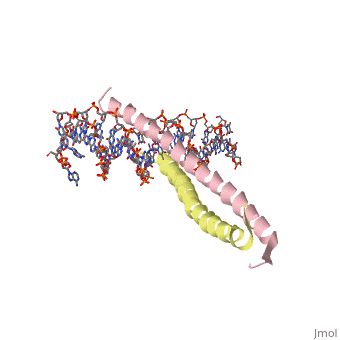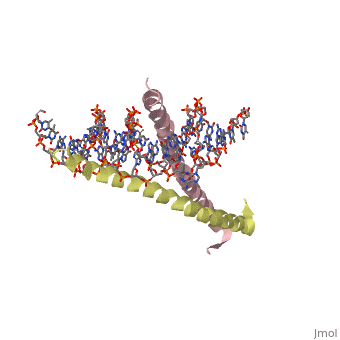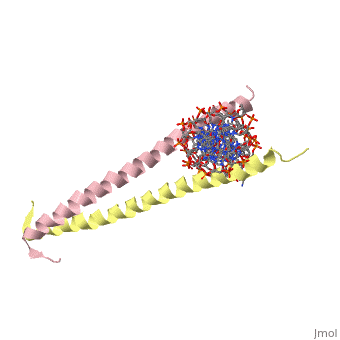
The yeast transcriptional activator GCN4 is 1 of over 30 identified eukaryotic proteins containing the basic region leucine zipper (bZIP) DNA-binding motif. We have determined the crystal structure of the GCN4 bZIP element complexed with DNA at 2.9 A resolution. The bZIP dimer is a pair of continuous alpha helices that form a parallel coiled coil over their carboxy-terminal 30 residues and gradually diverge toward their amino termini to pass through the major groove of the DNA-binding site. The coiled-coil dimerization interface is oriented almost perpendicular to the DNA axis, giving the complex the appearance of the letter T. There are no kinks or sharp bends in either bZIP monomer. Numerous contacts to DNA bases and phosphate oxygens are made by basic region residues that are conserved in the bZIP protein family. The details of the bZIP dimer interaction with DNA can explain recognition of the AP-1 site by the GCN4 protein. | |||
The GCN4 basic region leucine zipper binds DNA as a dimer of uninterrupted alpha helices: crystal structure of the protein-DNA complex.,Ellenberger TE, Brandl CJ, Struhl K, Harrison SC Cell. 1992 Dec 24;71(7):1223-37. PMID:1473154<ref>PMID:1473154</ref> | |||
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |||
</div> | |||
==See Also== | |||
*[[Gcn4|Gcn4]] | |||
== | *[[User:John S. de Banzie/DNA-BindingLeucineZipper|User:John S. de Banzie/DNA-BindingLeucineZipper]] | ||
[[ | == References == | ||
<references/> | |||
== | __TOC__ | ||
< | </StructureSection> | ||
[[Category: Saccharomyces cerevisiae]] | [[Category: Saccharomyces cerevisiae]] | ||
[[Category: Brandl, C J.]] | [[Category: Brandl, C J.]] | ||
Revision as of 00:57, 30 September 2014
THE GCN4 BASIC REGION LEUCINE ZIPPER BINDS DNA AS A DIMER OF UNINTERRUPTED ALPHA HELICES: CRYSTAL STRUCTURE OF THE PROTEIN-DNA COMPLEXTHE GCN4 BASIC REGION LEUCINE ZIPPER BINDS DNA AS A DIMER OF UNINTERRUPTED ALPHA HELICES: CRYSTAL STRUCTURE OF THE PROTEIN-DNA COMPLEX
Structural highlights
Evolutionary Conservation Check, as determined by ConSurfDB. You may read the explanation of the method and the full data available from ConSurf. Publication Abstract from PubMedThe yeast transcriptional activator GCN4 is 1 of over 30 identified eukaryotic proteins containing the basic region leucine zipper (bZIP) DNA-binding motif. We have determined the crystal structure of the GCN4 bZIP element complexed with DNA at 2.9 A resolution. The bZIP dimer is a pair of continuous alpha helices that form a parallel coiled coil over their carboxy-terminal 30 residues and gradually diverge toward their amino termini to pass through the major groove of the DNA-binding site. The coiled-coil dimerization interface is oriented almost perpendicular to the DNA axis, giving the complex the appearance of the letter T. There are no kinks or sharp bends in either bZIP monomer. Numerous contacts to DNA bases and phosphate oxygens are made by basic region residues that are conserved in the bZIP protein family. The details of the bZIP dimer interaction with DNA can explain recognition of the AP-1 site by the GCN4 protein. The GCN4 basic region leucine zipper binds DNA as a dimer of uninterrupted alpha helices: crystal structure of the protein-DNA complex.,Ellenberger TE, Brandl CJ, Struhl K, Harrison SC Cell. 1992 Dec 24;71(7):1223-37. PMID:1473154[1] From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine. See AlsoReferences |
| ||||||||||||||