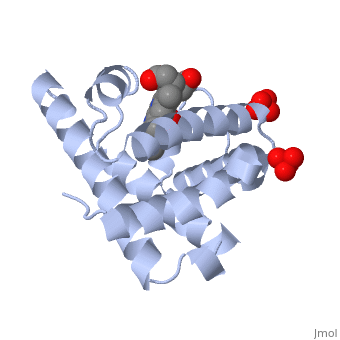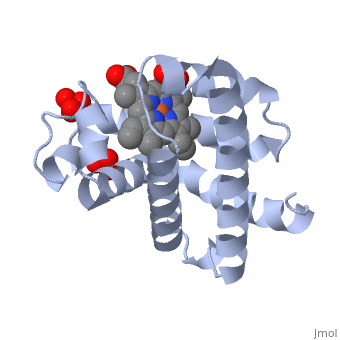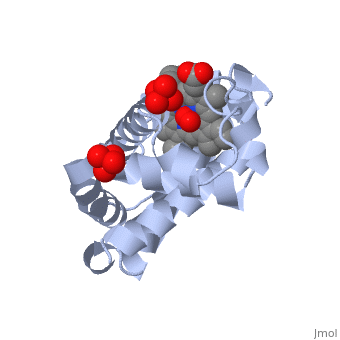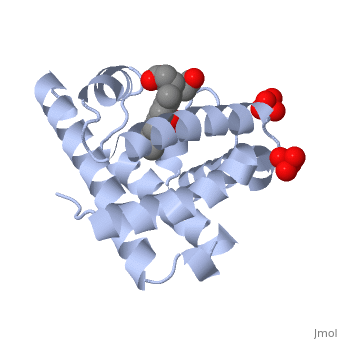Myoglobin
|
Caution: The text in this article and has not been updated to reflect what is actually available on this page. There is no zoom slider and no animate button. These were formerly present when an earlier version of Proteopedia supported Kinemages. A volunteer is needed to clean up and improve this article on a pedagogically and historically important protein. Green links are needed! Text referring to the now absent Kinemage is in gray. Eric Martz 01:09, 13 September 2014 (IDT) |
Myoglobin is a globular protein whose function is to store molecular oxygen. The fold of the protein is conserved but the sequence is more . The globin consists mostly of alpha helices shown in ; it has no beta sheets and its nonhelical segments mostly serve as links that connect the helices. Look down the barrel of some of the longer helices. Are they all straight? The eight structurally conserved alpha helices are labelled . The protein is colored as a N-->C rainbow in this view; the N terminus is blue, while the C terminus is red. The heme is shown in spacefilling form with its N, O, and Fe atoms displayed as blue, red, and orange balls. Note how the heme is almost completely enclosed by the globin. Which few chemical groups of the Heme are exposed to the solvent? (holding the mouse over the atoms reveals their identity.) Can you rationalize this exposure? View1 is the standard view of Mb. Rotate the protein around to convince yourself that the globin is approximately disc-shaped with a diameter that is about twice its thickness. Turn on the "Main Chain" button to display the polypeptide backbone in white with its N and O atoms represented by blue and red balls. How closely does the ribbon follow the main chain? Click "Animate" until the Mb ribbon is gray. Turn off the "Mb Ribbon" button. What are the orientations of the main chain carbonyl groups and the amide N atoms relative to each other which allows formation of the H-bonds of the alpha helices (not drawn)? View2 is a closeup of the heme from the same direction as View1. Turn on the "HemeLigand" button to display, in cyan, the sidechain of His 93, the proximal His, liganding the heme's Fe(II) ion (white bond). The Fe(II) is also liganded in a square-planar array by the heme's four pyrrole N atoms and hence has a total of 5 pyramidally arranged ligands. In oxyMb, the reversibly bound O2 molecule ligands the Fe from the opposite side of the heme as does the proximal His so that the Fe(II) becomes octahedrally coordinated. The Fe atom is not oxidized by its O2 ligand; it remains in the Fe(II) oxidation state. Rotate the image about the vertical axis until you see heme edge-on. Is the Heme planar? Note that the Fe atom is displaced towards the proximal His by 0.55 Å from the best plane though the porphyrin ring atoms. In oxyMb, the Fe is only 0.22 Å out of the heme plane and still on the side of the proximal His (for details see Oxymyoglobin). See also Molecular Playground/Myoglobin Myoglobin page in Spanish Myoglobin-Physeter-catodon-structure.
|
| ||||||||||
Former exercise in large part by John H. Connor (present address: Department of Microbiology, Boston University School of Medicine, 850 Harrison Ave, Boston, MA, 02118, USA). Revised by Ann Taylor
3D Structures of Myoglobin3D Structures of Myoglobin
Updated on 29-March-2015
Myoglobin (Mb) is an oxygen binding protein found in muscle tissue. It contains a heme group. Metmyoglobin (MMb) is the oxidized form of myoglobin.
((3qm5, 3qm6 – btMb – blackfin tuna
- Mb mutants uncomplexed
- 3m38, 3m39, 3m3a, 3m3b, 3k9z, 3h57, 3h58, 2e2y, 2ef2, 2oh8, 2oh9, 2oha, 2ohb, 2blh, 2bli, 1h1x, 1co8, 1n9h, 1n9i, 1n9x, 1naz, 1f63, 1f65, 1dti, 1co9, 1cp0, 1cp5, 1cpw, 1ch1, 1ch2, 1ch3, 1ch5, 1ch7, 1ch9, 1cik, 1cio, 1ofk, 1ofj, 102m, 1obm, 2mbw, 1tes, 1irc, 1mti, 1mtj, 1mtk, 1mlf, 1mlg, 1mlh, 1mlj, 1mlk, 1mll, 1mlm, 1mlo, 1mlr, 1mln, 1mls, 2mga, 2mgb, 2mgc, 2mgd, 2mge, 2mgf, 2mgg, 2mgh, 2mgi, 2mgj, 2mgk, 2mgl, 2mgm, 2spl, 2spm, 2spn, 2spo, 1fcs, 1j52, 1lue, 1o16, 3ogb, 3obd, 3ock, 3sdn, 3o89, 4fwx, 4fwz, 4it8 - SwMb (mutant)
- 2bwh - SwMb (mutant) – Laue
- 3hc9, 3hen, 3heo, 3hep, 1nz2, 1nz3, 1rse, 1abs, 1xch, 1hrm, 1yma, 3rj6 - hoMb (mutant) – horse
- 2vlx , 2vly, 2vlz, 2vm0 – hoMb fragment
- 1dm1 – AlMb (mutant)
- 2mm1, 3rgk – hMb (mutant) – human
- 3m38, 3m39, 3m3a, 3m3b, 3k9z, 3h57, 3h58, 2e2y, 2ef2, 2oh8, 2oh9, 2oha, 2ohb, 2blh, 2bli, 1h1x, 1co8, 1n9h, 1n9i, 1n9x, 1naz, 1f63, 1f65, 1dti, 1co9, 1cp0, 1cp5, 1cpw, 1ch1, 1ch2, 1ch3, 1ch5, 1ch7, 1ch9, 1cik, 1cio, 1ofk, 1ofj, 102m, 1obm, 2mbw, 1tes, 1irc, 1mti, 1mtj, 1mtk, 1mlf, 1mlg, 1mlh, 1mlj, 1mlk, 1mll, 1mlm, 1mlo, 1mlr, 1mln, 1mls, 2mga, 2mgb, 2mgc, 2mgd, 2mge, 2mgf, 2mgg, 2mgh, 2mgi, 2mgj, 2mgk, 2mgl, 2mgm, 2spl, 2spm, 2spn, 2spo, 1fcs, 1j52, 1lue, 1o16, 3ogb, 3obd, 3ock, 3sdn, 3o89, 4fwx, 4fwz, 4it8 - SwMb (mutant)
- Mb containing a non-Fe protoporphyrin
- 1yog, 1yoh, 1yoi, 1myz – SwMb+Co protoporphyrin
- 3mn0 – SwMb (mutant)+Cu protoporphyrin+cyanide
- 4mxk, 4mxl - SwMb (mutant)+Zn protoporphyrin
- 1j3f – SwMb+Cr salophen
- 1ufj, 1ufp – SwMb (mutant)+Fe salophen
- 2o58, 2o5b, 3wi8 – hoMb+Mn protoporphyrin
- 2o5l - hoMb+Mn protoporphyrin +methanol
- 2o5m - hoMb+Mn protoporphyrin +azide
- 2o5o, 2o5q - hoMb+Mn protoporphyrin +NO2
- 2o5s - hoMb+Co protoporphyrin +NO2
- 2o5t - hoMb+Co protoporphyrin
- 3rjn - hoMb+Zn deuteroporphyrin
- 1yog, 1yoh, 1yoi, 1myz – SwMb+Co protoporphyrin
- Mb+NO
- Mb+NO2
- Mb+O2
- Mb+CO
- 2bw9, 1mz0 - SwMb (mutant)+CO – Laue
- 2g0r, 2g0s, 2g0v, 2g0x, 2g0z, 2g10, 2g11, 2g12, 2g14, 2blj, 1dxc, 1dxd, 1do1, 1do3, 1do4, 1do7, 1abs, 1mcy, 1mbc, 3nml – SwMb (mutant)+CO
- 1bzr , 1a6g, 1ajg, 1ajh, 1spe, 1mym, 1mbc – SwMb+CO
- 1myf - SwMb+CO – NMR
- 2mb5 - SwMb+CO – Neutron
- 1dwr, 1dws, 1dwt – hoMb+CO
- 1mdn, 1m6c – pMb (mutant)+CO
- 1mwc, 1yca – pMb+CO
- 3qm7 – btMb + CO
- 2bw9, 1mz0 - SwMb (mutant)+CO – Laue
- Mb+OH
- Mb+cyano compounds
- 2jho – SwMb cyanoMet
- 3qm8 - btMb cyanoMet
- 1ebc, 2cmm - SwMb+cyanide
- 1iop – SwMb+cyanide+hemin
- 108m, 101m, 104m, 105m, 2myc, 2myd, 2mya, 2mye, 2myb – SwMb+isocyanides
- 103m, 107m, 111m, 106m, 109m, 110m, 112m – SwMb (mutant) +isocyanides
- 3ba2 – hoMb+cyanide
- 2fal – AlMb+cyanide
- 2fam – AlMb+thiocyanate
- 1lht – stMb+cyanide
- 1emy – Mb+cyanide – asian elephant
- 2jho – SwMb cyanoMet
- Mb + N3
- 3qm9 – btMb + N3
- MMb
- Other complexes of Mb
- 3a2g – SwMb (mutant)+fluorescein
- 2w6x, 2w6y - SwMb (mutant)+Xe
- 2w6w - SwMb +Xe
- 2eb8, 2eb9 – SwMb+Cu
- 4fwy - SwMb (mutant)+ Cu
- 3ase – SwMb + RuO3
- 2ekt, 2eku – SwMb+methyl-depropionatehemin
- 2d6c – SwMb+porphycene
- 1swm, 1mbi – SwMb+imidazole
- 3u3e – SwMb + phenol
- 4h07 - SwMb (mutant)+ phenol
- 4h0b - SwMb (mutant)+ DMSO
- 1wvp – SwMb+tetrazolyl histidine
- 2evk – SwMb (mutant)+acetic acid
- 2evp - SwMb (mutant)+mercaptoethanol
- 1v9q - SwMb (mutant)+Mn salophen derivative
- 1dtm - SwMb (mutant)+4-methylimidazole
- 2nsr – hoMb+nitro methane
- 1npg – hoMb+nitrosoethane
- 2nss - hoMb+nitro benzene
- 1azi - hoMb+azide
- 1bje – hoMb (mutant)+azide
- 1nz4 – hoMb (mutant)+Cd
- 1nz5 - hoMb (mutant)+Mn
- 2in4 - hoMb+Zn-DME
- 1gjn – hoMb+hydroxide
- 2nrm – BtMb S-nitrosylated
- 5mba – AlMb+azide
- 1mni – pMb+imidazole
- 3qma - btMb+imidazole
- 3a2g – SwMb (mutant)+fluorescein
- apo-Mb