1tyf

| |||||||||
| 1tyf, resolution 2.30Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Activity: | Endopeptidase Clp, with EC number 3.4.21.92 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
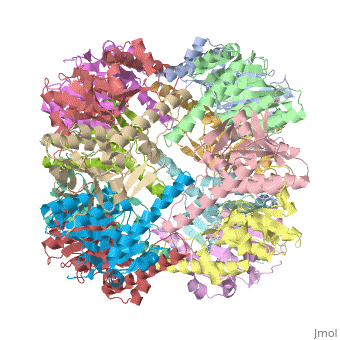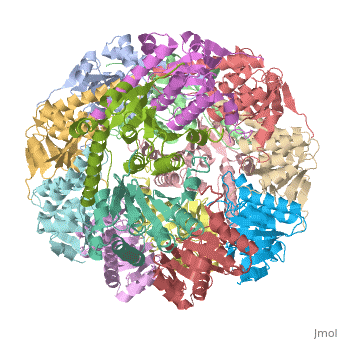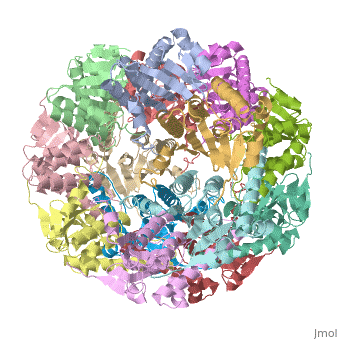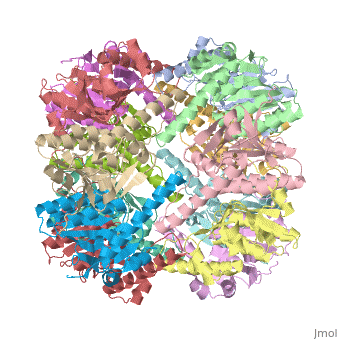
THE STRUCTURE OF CLPP AT 2.3 ANGSTROM RESOLUTION SUGGESTS A MODEL FOR ATP-DEPENDENT PROTEOLYSISTHE STRUCTURE OF CLPP AT 2.3 ANGSTROM RESOLUTION SUGGESTS A MODEL FOR ATP-DEPENDENT PROTEOLYSIS
We have determined the crystal structure of the proteolytic component of the caseinolytic Clp protease (ClpP) from E. coli at 2.3 A resolution using an ab initio phasing procedure that exploits the internal 14-fold symmetry of the oligomer. The structure of a ClpP monomer has a distinct fold that defines a fifth structural family of serine proteases but a conserved catalytic apparatus. The active protease resembles a hollow, solid-walled cylinder composed of two 7-fold symmetric rings stacked back-to-back. Its 14 proteolytic active sites are located within a central, roughly spherical chamber approximately 51 A in diameter. Access to the proteolytic chamber is controlled by two axial pores, each having a minimum diameter of approximately 10 A. From the structural features of ClpP, we suggest a model for its action in degrading proteins.
The structure of ClpP at 2.3 A resolution suggests a model for ATP-dependent proteolysis., Wang J, Hartling JA, Flanagan JM, Cell. 1997 Nov 14;91(4):447-56. PMID:9390554
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.
About this StructureAbout this Structure
1tyf is a 14 chain structure with sequence from Escherichia coli. Full crystallographic information is available from OCA.