1jb0
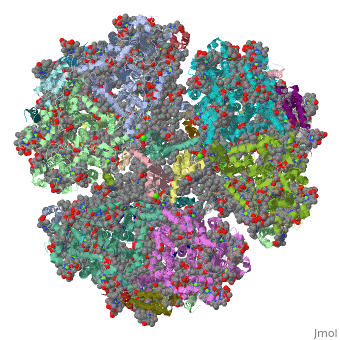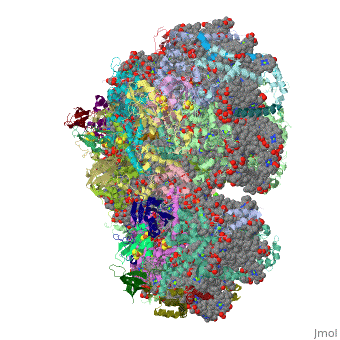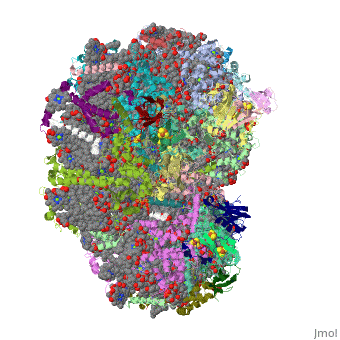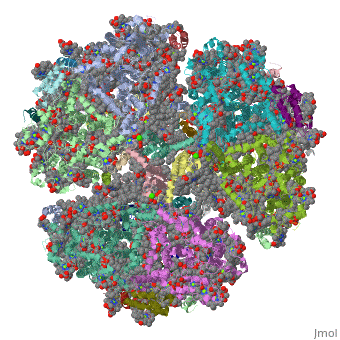
Crystal Structure of Photosystem I: a Photosynthetic Reaction Center and Core Antenna System from CyanobacteriaCrystal Structure of Photosystem I: a Photosynthetic Reaction Center and Core Antenna System from Cyanobacteria
Structural highlights
FunctionPSAA_THEVB PsaA and PsaB bind P700, the primary electron donor of photosystem I (PSI), as well as the electron acceptors A0, A1 and FX. PSI is a plastocyanin/cytochrome c6-ferredoxin oxidoreductase, converting photonic excitation into a charge separation, which transfers an electron from the donor P700 chlorophyll pair to the spectroscopically characterized acceptors A0, A1, FX, FA and FB in turn. Oxidized P700 is reduced on the lumenal side of the thylakoid membrane by plastocyanin or cytochrome c6.[1] [2] [3] [4] Evolutionary Conservation Check, as determined by ConSurfDB. You may read the explanation of the method and the full data available from ConSurf. Publication Abstract from PubMedLife on Earth depends on photosynthesis, the conversion of light energy from the Sun to chemical energy. In plants, green algae and cyanobacteria, this process is driven by the cooperation of two large protein-cofactor complexes, photosystems I and II, which are located in the thylakoid photosynthetic membranes. The crystal structure of photosystem I from the thermophilic cyanobacterium Synechococcus elongatus described here provides a picture at atomic detail of 12 protein subunits and 127 cofactors comprising 96 chlorophylls, 2 phylloquinones, 3 Fe4S4 clusters, 22 carotenoids, 4 lipids, a putative Ca2+ ion and 201 water molecules. The structural information on the proteins and cofactors and their interactions provides a basis for understanding how the high efficiency of photosystem I in light capturing and electron transfer is achieved. Three-dimensional structure of cyanobacterial photosystem I at 2.5 A resolution.,Jordan P, Fromme P, Witt HT, Klukas O, Saenger W, Krauss N Nature. 2001 Jun 21;411(6840):909-17. PMID:11418848[5] From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine. See AlsoReferences
|
| ||||||||||||||||||