1aoi: Difference between revisions
No edit summary |
No edit summary |
||
| Line 3: | Line 3: | ||
<StructureSection load='1aoi' size='340' side='right'caption='[[1aoi]], [[Resolution|resolution]] 2.80Å' scene=''> | <StructureSection load='1aoi' size='340' side='right'caption='[[1aoi]], [[Resolution|resolution]] 2.80Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
<table><tr><td colspan='2'>[[1aoi]] is a 10 chain structure. The July 2000 RCSB PDB [ | <table><tr><td colspan='2'>[[1aoi]] is a 10 chain structure. The July 2000 RCSB PDB [https://pdb.rcsb.org/pdb/static.do?p=education_discussion/molecule_of_the_month/index.html Molecule of the Month] feature on ''Nucleosome'' by David S. Goodsell is [https://dx.doi.org/10.2210/rcsb_pdb/mom_2000_7 10.2210/rcsb_pdb/mom_2000_7]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1AOI OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1AOI FirstGlance]. <br> | ||
</td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=MN:MANGANESE+(II)+ION'>MN</scene></td></tr> | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=MN:MANGANESE+(II)+ION'>MN</scene></td></tr> | ||
<tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1aoi FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1aoi OCA], [https://pdbe.org/1aoi PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1aoi RCSB], [https://www.ebi.ac.uk/pdbsum/1aoi PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1aoi ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
[[ | [[https://www.uniprot.org/uniprot/H2A1_XENLA H2A1_XENLA]] Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. [[https://www.uniprot.org/uniprot/H4_XENLA H4_XENLA]] Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. [[https://www.uniprot.org/uniprot/H32_XENLA H32_XENLA]] Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. [[https://www.uniprot.org/uniprot/H2B11_XENLA H2B11_XENLA]] Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. | ||
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 30: | Line 30: | ||
==See Also== | ==See Also== | ||
*[[Histone|Histone | *[[Histone 3D structures|Histone 3D structures]] | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 13:22, 17 February 2021
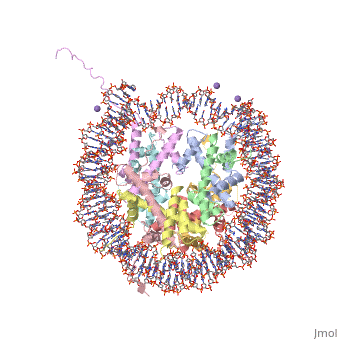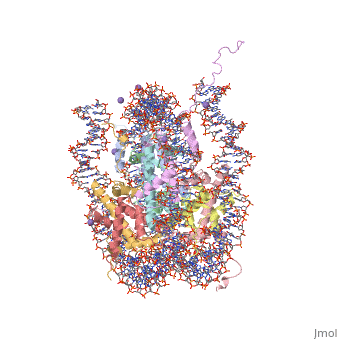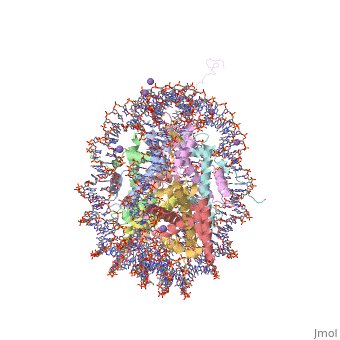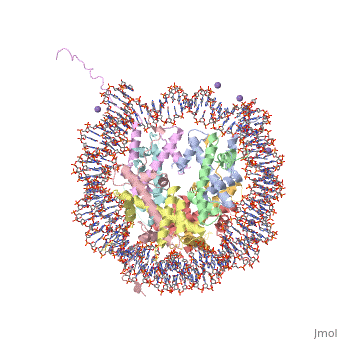
COMPLEX BETWEEN NUCLEOSOME CORE PARTICLE (H3,H4,H2A,H2B) AND 146 BP LONG DNA FRAGMENTCOMPLEX BETWEEN NUCLEOSOME CORE PARTICLE (H3,H4,H2A,H2B) AND 146 BP LONG DNA FRAGMENT
Structural highlights
Function[H2A1_XENLA] Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. [H4_XENLA] Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. [H32_XENLA] Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. [H2B11_XENLA] Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. Evolutionary Conservation Check, as determined by ConSurfDB. You may read the explanation of the method and the full data available from ConSurf. Publication Abstract from PubMedThe X-ray crystal structure of the nucleosome core particle of chromatin shows in atomic detail how the histone protein octamer is assembled and how 146 base pairs of DNA are organized into a superhelix around it. Both histone/histone and histone/DNA interactions depend on the histone fold domains and additional, well ordered structure elements extending from this motif. Histone amino-terminal tails pass over and between the gyres of the DNA superhelix to contact neighbouring particles. The lack of uniformity between multiple histone/DNA-binding sites causes the DNA to deviate from ideal superhelix geometry. Crystal structure of the nucleosome core particle at 2.8 A resolution.,Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ Nature. 1997 Sep 18;389(6648):251-60. PMID:9305837[1] From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine. See AlsoReferences
|
| ||||||||||||||||