2h24: Difference between revisions
No edit summary |
No edit summary |
||
| Line 7: | Line 7: | ||
|ACTIVITY= | |ACTIVITY= | ||
|GENE= | |GENE= | ||
|DOMAIN= | |||
|RELATEDENTRY=[[1inr|1INR]] | |||
|RESOURCES=<span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2h24 FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2h24 OCA], [http://www.ebi.ac.uk/pdbsum/2h24 PDBsum], [http://www.rcsb.org/pdb/explore.do?structureId=2h24 RCSB]</span> | |||
}} | }} | ||
| Line 14: | Line 17: | ||
==Overview== | ==Overview== | ||
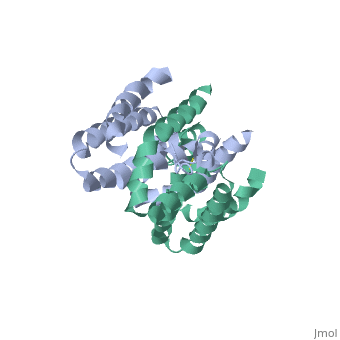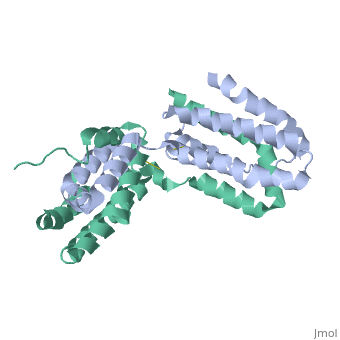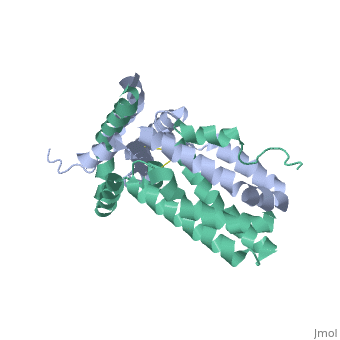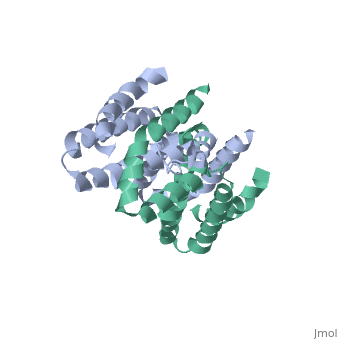
Interleukin-10 receptor 2 (IL-10R2) is a critical component of the IL-10.IL-10R1.IL-10R2 complex which regulates IL-10-mediated immunomodulatory responses. The ternary IL-10 signaling complex is assembled in a sequential order with the IL-10.IL-10R1 interaction occurring first followed by engagement of the IL-10R2 chain. In this study we map the IL-10R2 binding site on IL-10 using surface plasmon resonance and cell-based assays. Critical IL-10R2 binding residues are located in helix A adjacent to the previously identified IL-10R1 recognition surface. Interestingly, IL-10R2 binding residues located in the N-terminal end of helix A exhibit large structural differences between unbound cIL-10 and cIL-10.IL-10R1 crystal structures. This suggests IL-10R1-induced conformational changes regulate IL-10R2 binding and assembly of the ternary IL-10.IL-10R1.IL-10R2 complex. The basic mechanistic features of the assembly process are likely shared by six additional class-2 cytokines (viral IL-10s, IL-22, IL-26, IL-28A, IL28B, and IL-29) to promote IL-10R2 binding to six additional receptor complexes. These studies highlight the importance of structure in regulating low affinity protein-protein interactions and IL-10 signal transduction. | Interleukin-10 receptor 2 (IL-10R2) is a critical component of the IL-10.IL-10R1.IL-10R2 complex which regulates IL-10-mediated immunomodulatory responses. The ternary IL-10 signaling complex is assembled in a sequential order with the IL-10.IL-10R1 interaction occurring first followed by engagement of the IL-10R2 chain. In this study we map the IL-10R2 binding site on IL-10 using surface plasmon resonance and cell-based assays. Critical IL-10R2 binding residues are located in helix A adjacent to the previously identified IL-10R1 recognition surface. Interestingly, IL-10R2 binding residues located in the N-terminal end of helix A exhibit large structural differences between unbound cIL-10 and cIL-10.IL-10R1 crystal structures. This suggests IL-10R1-induced conformational changes regulate IL-10R2 binding and assembly of the ternary IL-10.IL-10R1.IL-10R2 complex. The basic mechanistic features of the assembly process are likely shared by six additional class-2 cytokines (viral IL-10s, IL-22, IL-26, IL-28A, IL28B, and IL-29) to promote IL-10R2 binding to six additional receptor complexes. These studies highlight the importance of structure in regulating low affinity protein-protein interactions and IL-10 signal transduction. | ||
==About this Structure== | ==About this Structure== | ||
| Line 29: | Line 29: | ||
[[Category: alpha-helix bundle]] | [[Category: alpha-helix bundle]] | ||
''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Mon Mar 31 03:24:24 2008'' | ||
Revision as of 03:24, 31 March 2008
| |||||||
| , resolution 2.0Å | |||||||
|---|---|---|---|---|---|---|---|
| Related: | 1INR
| ||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
Crystal structure of human IL-10
OverviewOverview
Interleukin-10 receptor 2 (IL-10R2) is a critical component of the IL-10.IL-10R1.IL-10R2 complex which regulates IL-10-mediated immunomodulatory responses. The ternary IL-10 signaling complex is assembled in a sequential order with the IL-10.IL-10R1 interaction occurring first followed by engagement of the IL-10R2 chain. In this study we map the IL-10R2 binding site on IL-10 using surface plasmon resonance and cell-based assays. Critical IL-10R2 binding residues are located in helix A adjacent to the previously identified IL-10R1 recognition surface. Interestingly, IL-10R2 binding residues located in the N-terminal end of helix A exhibit large structural differences between unbound cIL-10 and cIL-10.IL-10R1 crystal structures. This suggests IL-10R1-induced conformational changes regulate IL-10R2 binding and assembly of the ternary IL-10.IL-10R1.IL-10R2 complex. The basic mechanistic features of the assembly process are likely shared by six additional class-2 cytokines (viral IL-10s, IL-22, IL-26, IL-28A, IL28B, and IL-29) to promote IL-10R2 binding to six additional receptor complexes. These studies highlight the importance of structure in regulating low affinity protein-protein interactions and IL-10 signal transduction.
About this StructureAbout this Structure
2H24 is a Single protein structure of sequence from Homo sapiens. Full crystallographic information is available from OCA.
ReferenceReference
Conformational changes mediate interleukin-10 receptor 2 (IL-10R2) binding to IL-10 and assembly of the signaling complex., Yoon SI, Logsdon NJ, Sheikh F, Donnelly RP, Walter MR, J Biol Chem. 2006 Nov 17;281(46):35088-96. Epub 2006 Sep 18. PMID:16982608
Page seeded by OCA on Mon Mar 31 03:24:24 2008