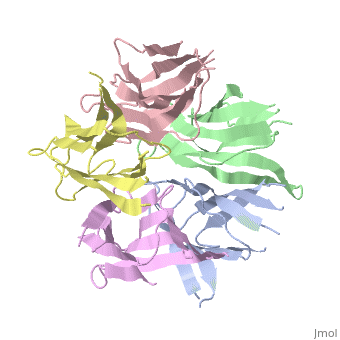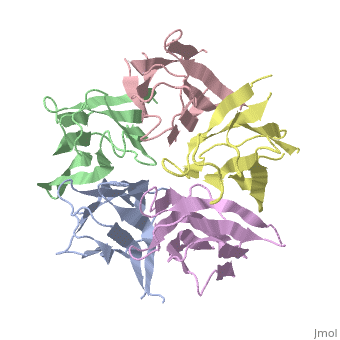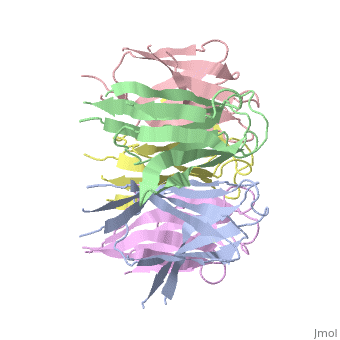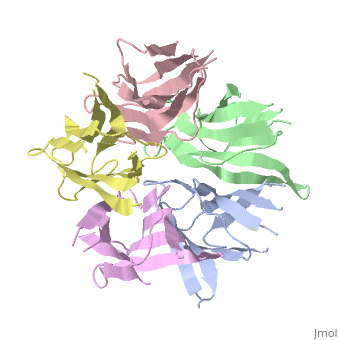1k5j: Difference between revisions
No edit summary |
No edit summary |
||
| Line 16: | Line 16: | ||
<text>to colour the structure by Evolutionary Conservation</text> | <text>to colour the structure by Evolutionary Conservation</text> | ||
</jmolCheckbox> | </jmolCheckbox> | ||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/ | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1k5j ConSurf]. | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
<div style="background-color:#fffaf0;"> | <div style="background-color:#fffaf0;"> | ||
Revision as of 03:09, 10 February 2016
The Crystal Structure of Nucleoplasmin-CoreThe Crystal Structure of Nucleoplasmin-Core
Structural highlights
Function[NUPL_XENLA] Core histones chaperone involved in chromatin reprogramming, specially during fertilization and early embryonic development. Nucleoplasmin is an acidic protein which is able to assemble nucleosomes by binding histones and transferring them to DNA. Evolutionary Conservation Check, as determined by ConSurfDB. You may read the explanation of the method and the full data available from ConSurf. Publication Abstract from PubMedThe efficient assembly of histone complexes and nucleosomes requires the participation of molecular chaperones. Currently, there is a paucity of data on their mechanism of action. We now present the structure of an N-terminal domain of nucleoplasmin (Np-core) at 2.3 A resolution. The Np-core monomer is an eight-stranded beta barrel that fits snugly within a stable pentamer. In the crystal, two pentamers associate to form a decamer. We show that both Np and Np-core are competent to assemble large complexes that contain the four core histones. Further experiments and modeling suggest that these complexes each contain five histone octamers which dock to a central Np decamer. This work has important ramifications for models of histone storage, sperm chromatin decondensation, and nucleosome assembly. The crystal structure of nucleoplasmin-core: implications for histone binding and nucleosome assembly.,Dutta S, Akey IV, Dingwall C, Hartman KL, Laue T, Nolte RT, Head JF, Akey CW Mol Cell. 2001 Oct;8(4):841-53. PMID:11684019[1] From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine. See AlsoReferences |
| ||||||||||||||||