Tropomyosin: Difference between revisions
Michal Harel (talk | contribs) No edit summary |
Michal Harel (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
<StructureSection load='' size='350' side='right' scene='User:Gregory_Hoeprich/Sandbox_1/Tropomyosin_dimer/2' caption='Pig tropomyosin (PDB code [[1c1g]])'> | <StructureSection load='' size='350' side='right' scene='User:Gregory_Hoeprich/Sandbox_1/Tropomyosin_dimer/2' caption='Pig tropomyosin (PDB code [[1c1g]])'> | ||
[[Image:B Lehman1.jpg | thumb | 300 x 430px | left | alt text | '''Tropomyosin''' (seen in yellow and red) wrapped around actin filaments, which are EM reconstructions with G-actin ribbion structures filling in the EM structure. (Picture generated from William Lehman's [http://www.bumc.bu.edu/phys-biophys/research/filhel/ Website]) ]] | [[Image:B Lehman1.jpg | thumb | 300 x 430px | left | alt text | '''Tropomyosin''' (seen in yellow and red) wrapped around actin filaments, which are EM reconstructions with G-actin ribbion structures filling in the EM structure. (Picture generated from William Lehman's [http://www.bumc.bu.edu/phys-biophys/research/filhel/ Website]) ]] | ||
__TOC__ | |||
==Function== | ==Function== | ||
'''[[Tropomyosin]] (TM)''' is an [[actin]] binding protein, which consists of a coiled-coil dimer (see left) and forms a polymer along the length of actin by a head-to-tail overlap along the major grove of actin (see down & left)<ref name="Gunning">Tropomyosins. I. Gunning, Peter, 1950- II. Series.[DNLM: 1. Tropomyosin. W1 AD559 v.644 2008 / WE 500 T856 2008]</ref>. See [[Coiled coil]]. The head-to-tail overlap allows flexibility between the tropomyosin dimers so it will lay unstrained along the filament<ref name="Gunning"/>. Each tropomyosin molecule spans seven actin monomers within a filament and lays N- to C- terminally from actin's pointed to barbed end<ref name="Frye">PMID:20465283</ref>. The 284 amino acid helix has a length of 420 Angstroms and has a molecular weight around 65-70 kilodaltons (vertebrate tropomyosin)<ref name="Gunning"/><ref name="Whitby">PMID:10651038</ref>. A few of tropomyosin's characteristics as an actin binding protein includes regulation, stabilization and recruitment. | '''[[Tropomyosin]] (TM)''' is an [[actin]] binding protein, which consists of a coiled-coil dimer (see left) and forms a polymer along the length of actin by a head-to-tail overlap along the major grove of actin (see down & left)<ref name="Gunning">Tropomyosins. I. Gunning, Peter, 1950- II. Series.[DNLM: 1. Tropomyosin. W1 AD559 v.644 2008 / WE 500 T856 2008]</ref>. See [[Coiled coil]]. The head-to-tail overlap allows flexibility between the tropomyosin dimers so it will lay unstrained along the filament<ref name="Gunning"/>. Each tropomyosin molecule spans seven actin monomers within a filament and lays N- to C- terminally from actin's pointed to barbed end<ref name="Frye">PMID:20465283</ref>. The 284 amino acid helix has a length of 420 Angstroms and has a molecular weight around 65-70 kilodaltons (vertebrate tropomyosin)<ref name="Gunning"/><ref name="Whitby">PMID:10651038</ref>. A few of tropomyosin's characteristics as an actin binding protein includes regulation, stabilization and recruitment. | ||
| Line 30: | Line 31: | ||
<br /> | <br /> | ||
== Post-Translational Modifications == | |||
There are two types of post-translational modifications to tropomyosin: phosphorylation acetylation<ref name="Gunning"/>. Phosphorylation occurs on amino acid <scene name='41/410306/Tropomyosin_dimer_ser229/1'>Ser-229</scene><ref name="Gunning"/>. This phosphorylation is occurs as a result of oxidative stress, which is associated with actin remodeling and recruitment of additional tropomyosin into stress fibers<ref name="Gunning"/>. The acetylation occurs on the N-terminus of the N-terminal methionine, which is essential for: coiled-coil stability, overlap formation and actin binding<ref name="Frye"/><ref name="Gunning"/>. | There are two types of post-translational modifications to tropomyosin: phosphorylation acetylation<ref name="Gunning"/>. Phosphorylation occurs on amino acid <scene name='41/410306/Tropomyosin_dimer_ser229/1'>Ser-229</scene><ref name="Gunning"/>. This phosphorylation is occurs as a result of oxidative stress, which is associated with actin remodeling and recruitment of additional tropomyosin into stress fibers<ref name="Gunning"/>. The acetylation occurs on the N-terminus of the N-terminal methionine, which is essential for: coiled-coil stability, overlap formation and actin binding<ref name="Frye"/><ref name="Gunning"/>. | ||
=== Evolutionary Conservation === | === Evolutionary Conservation === | ||
Tropomyosin is highly conserved actin binding protein, which is found in Eukarya from the animal kingdom to yeast, with the exception of plants<ref name="Gunning"/>. The earliest characterization of tropomyosin gene lineage was in yeast (budding and fission yeast)<ref name="Gunning"/><ref name="Drees">PMID:7844152</ref>. These genes are known as [http://en.wikipedia.org/wiki/TPM1 TPM1] and [http://en.wikipedia.org/wiki/TPM2 TPM2], respectively, and share 64.5% sequence identity<ref name="Gunning"/><ref name="Drees"/>. As we move away from unicellular organisms and into multicellular invertebrates, tropomyosin genes in nematodes slightly diverge from yeast, but have 85-90% sequence identity between their genes<ref name="Gunning"/>. Further analysis for vertebrates show there are four genes that generate over 40 known mammalian isoforms of tropomyosin, which are synthesized by exon splicing<ref name="Gunning"/>. The slight evolution change of tropomyosin has occurred as a result of the increasing need of tropomyosin to function in different systems, but tropomyosin has evolutionarily stayed well conserved because of the basic structural pressures imposed on the protein<ref name="Gunning"/>. It is interesting to note, the region with the least conservation has been in the N and C terminus. This is the result of head-tail interactions changing to accommodate different polymer confirmations along actin for different functions<ref name="Gunning"/>. (To see this evolutionary conservation, go to the top right image of the web page and click on the "show" link, which is to the right of "Evolutionary Conservation".) | Tropomyosin is highly conserved actin binding protein, which is found in Eukarya from the animal kingdom to yeast, with the exception of plants<ref name="Gunning"/>. The earliest characterization of tropomyosin gene lineage was in yeast (budding and fission yeast)<ref name="Gunning"/><ref name="Drees">PMID:7844152</ref>. These genes are known as [http://en.wikipedia.org/wiki/TPM1 TPM1] and [http://en.wikipedia.org/wiki/TPM2 TPM2], respectively, and share 64.5% sequence identity<ref name="Gunning"/><ref name="Drees"/>. As we move away from unicellular organisms and into multicellular invertebrates, tropomyosin genes in nematodes slightly diverge from yeast, but have 85-90% sequence identity between their genes<ref name="Gunning"/>. Further analysis for vertebrates show there are four genes that generate over 40 known mammalian isoforms of tropomyosin, which are synthesized by exon splicing<ref name="Gunning"/>. The slight evolution change of tropomyosin has occurred as a result of the increasing need of tropomyosin to function in different systems, but tropomyosin has evolutionarily stayed well conserved because of the basic structural pressures imposed on the protein<ref name="Gunning"/>. It is interesting to note, the region with the least conservation has been in the N and C terminus. This is the result of head-tail interactions changing to accommodate different polymer confirmations along actin for different functions<ref name="Gunning"/>. (To see this evolutionary conservation, go to the top right image of the web page and click on the "show" link, which is to the right of "Evolutionary Conservation".) | ||
== Tropomyosin's Function: Muscle and Non-Muscle Systems == | |||
Muscle tissue is comprised of many muscle fibers or cells. Those muscle fibers consist of myofibrils that contain a series of contractile units called sarcomeres. Within this unit contains thick filaments, comprised mainly of [[myosin]], and thin filaments, which contain [[actin]], tropomyosin and [[troponin]]. Tropomyosin in striated muscle systems (skeletal and cardiac) acts to inhibit the myosin cross-bridges from binding to the myosin binding site on thin filaments, this tropomyosin state is in the "Blocked" position<ref name="Lehman">PMID:19341744</ref>. When the muscle is stimulated, there is a rise in intracellular calcium stemming from a cascade of cellular processes. As the calcium is bathing the sarcomere, it will bind to the troponin complex, which is bound to both actin and tropomyosin<ref name="Tyska">PMID:11810692</ref>. The troponin will displace the tropomyosin from a "Blocked" to a "Closed" position<ref name="Lehman"/>. This transition allows the myosin head to interact weakly with the myosin binding site<ref name="Tyska"/>. The tropomyosin is displaced to its final position, "Open" state, along actin filament as myosin binds to its site<ref name="Lehman"/>. These three tropomyosin states along the filament is referred to as the three state model<ref name="Lehman"/>. As the intracellular calcium concentration falls, the troponin no longer is able to displace tropomyosin and it will transition back to the "Blocked" state<ref name="Lehman"/>. | Muscle tissue is comprised of many muscle fibers or cells. Those muscle fibers consist of myofibrils that contain a series of contractile units called sarcomeres. Within this unit contains thick filaments, comprised mainly of [[myosin]], and thin filaments, which contain [[actin]], tropomyosin and [[troponin]]. Tropomyosin in striated muscle systems (skeletal and cardiac) acts to inhibit the myosin cross-bridges from binding to the myosin binding site on thin filaments, this tropomyosin state is in the "Blocked" position<ref name="Lehman">PMID:19341744</ref>. When the muscle is stimulated, there is a rise in intracellular calcium stemming from a cascade of cellular processes. As the calcium is bathing the sarcomere, it will bind to the troponin complex, which is bound to both actin and tropomyosin<ref name="Tyska">PMID:11810692</ref>. The troponin will displace the tropomyosin from a "Blocked" to a "Closed" position<ref name="Lehman"/>. This transition allows the myosin head to interact weakly with the myosin binding site<ref name="Tyska"/>. The tropomyosin is displaced to its final position, "Open" state, along actin filament as myosin binds to its site<ref name="Lehman"/>. These three tropomyosin states along the filament is referred to as the three state model<ref name="Lehman"/>. As the intracellular calcium concentration falls, the troponin no longer is able to displace tropomyosin and it will transition back to the "Blocked" state<ref name="Lehman"/>. | ||
<br /> | <br /> | ||
<br /> | <br /> | ||
In non-muscle systems, as mentioned above, tropomyosin performs a wide variety of functions inside of cells. One of the specific functions tropomyosin performs within cells is recruitment of different myosin motors. Clayton ''et al'', 2010 showed actin decorated with tropomyosin increased myosin-V (cargo transporter) motility as opposed to bare actin<ref name="Clayton">PMID:20705471</ref>. It was also shown that myosin-I (cargo transporter and tension senor) lost its motility function on actin decorated with tropomyosin<ref name="Clayton"/>. Further evidence from a related paper, Stark ''et al'' 2010, shows tropomyosin increases the affinity of myosin-II to actin filaments<ref name="Stark">PMID:20110347</ref>. So actin decorated with tropomyosin appears to spatially regulate motor activity and thus cellular function. | In non-muscle systems, as mentioned above, tropomyosin performs a wide variety of functions inside of cells. One of the specific functions tropomyosin performs within cells is recruitment of different myosin motors. Clayton ''et al'', 2010 showed actin decorated with tropomyosin increased myosin-V (cargo transporter) motility as opposed to bare actin<ref name="Clayton">PMID:20705471</ref>. It was also shown that myosin-I (cargo transporter and tension senor) lost its motility function on actin decorated with tropomyosin<ref name="Clayton"/>. Further evidence from a related paper, Stark ''et al'' 2010, shows tropomyosin increases the affinity of myosin-II to actin filaments<ref name="Stark">PMID:20110347</ref>. So actin decorated with tropomyosin appears to spatially regulate motor activity and thus cellular function. | ||
== 3D structures of Tropomyosin == | |||
[[Tropomyosin 3D structures]] | |||
</StructureSection> | </StructureSection> | ||
| Line 61: | Line 66: | ||
**[[3u1a]], [[3u1c]] – cTPM α-1 N terminal<br /> | **[[3u1a]], [[3u1c]] – cTPM α-1 N terminal<br /> | ||
**[[3u59]] – cTPM β N terminal<br /> | **[[3u59]] – cTPM β N terminal<br /> | ||
**[[7bjg]], [[7bjn]] – DmTPM 270-334 – ''Drosophila melanogaster''<br /> | |||
**[[2tma]] – TPM - model<br /> | **[[2tma]] – TPM - model<br /> | ||
| Line 70: | Line 76: | ||
**[[6g2t]], [[6cxj]], [[6cxi]] - hTPM + myosin-binding protein + actin – Cryo EM<br /> | **[[6g2t]], [[6cxj]], [[6cxi]] - hTPM + myosin-binding protein + actin – Cryo EM<br /> | ||
**[[7jh7]] - pTPM + myosin-7 + actin – Cryo EM<br /> | **[[7jh7]] - pTPM + myosin-7 + actin – Cryo EM<br /> | ||
**[[7ko4]], [[7kor5], [[7ko7]], [[7kon]], [[7kor]] - pTPM α-1 + troponin + actin – Cryo EM<br /> | |||
**[[5nog]], [[5noj]], [[5nol]] - pTPM α-1 + actin – Cryo EM<br /> | **[[5nog]], [[5noj]], [[5nol]] - pTPM α-1 + actin – Cryo EM<br /> | ||
**[[2efr]], [[2efs]], [[2d3e]] - raTPM α-1 C-terminal+GNC4 leucine zipper – rabbit<br /> | **[[2efr]], [[2efs]], [[2d3e]] - raTPM α-1 C-terminal+GNC4 leucine zipper – rabbit<br /> | ||
**[[4a7f]], [[4a7h]], [[4a7l]] - raTPM α-1 + myosin IE heavy chain + actin – Cryo EM<br /> | **[[4a7f]], [[4a7h]], [[4a7l]] - raTPM α-1 + myosin IE heavy chain + actin – Cryo EM<br /> | ||
**[[3j4k]] - raTPM + actin – Cryo EM<br /> | **[[3j4k]] - raTPM + actin – Cryo EM<br /> | ||
**[[7nep]] - mTPM α-1 + myosin-4 + myosin light chain + actin – Cryo EM<br /> | |||
**[[7qin]] - mTPM α-1 in actomyosin complex – Cryo EM<br /> | |||
**[[3j8a]], [[5jlf]] - mTPM + actin - Cryo EM<br /> | **[[3j8a]], [[5jlf]] - mTPM + actin - Cryo EM<br /> | ||
**[[2g9j]] - rTPM α-1 TM9A+GNC4 <br /> | **[[2g9j]] - rTPM α-1 TM9A+GNC4 <br /> | ||
**[[2w49]], [[2w4u]] – cTnnC+cTnnT+cTnnI+cTPM α-1 + cActin<br /> | **[[2w49]], [[2w4u]] – cTnnC+cTnnT+cTnnI+cTPM α-1 + cActin<br /> | ||
**[[7bjs]] – DmTPM 252-334 + kinesin heavy chain<br /> | |||
**[[2z5h]] – yTPM α-1 N-terminal+C-terminal+GNC4 leucine zipper+TnnT – yeast<br /> | **[[2z5h]] – yTPM α-1 N-terminal+C-terminal+GNC4 leucine zipper+TnnT – yeast<br /> | ||
**[[2z5i]] - yTPM α-1 N-terminal+C-terminal+GNC4 leucine zipper<br /> | **[[2z5i]] - yTPM α-1 N-terminal+C-terminal+GNC4 leucine zipper<br /> | ||
Revision as of 12:42, 18 May 2022
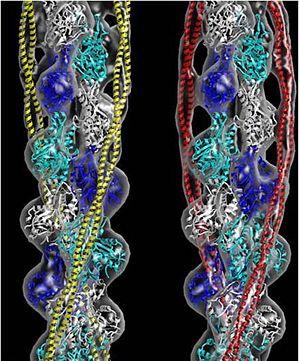 FunctionTropomyosin (TM) is an actin binding protein, which consists of a coiled-coil dimer (see left) and forms a polymer along the length of actin by a head-to-tail overlap along the major grove of actin (see down & left)[1]. See Coiled coil. The head-to-tail overlap allows flexibility between the tropomyosin dimers so it will lay unstrained along the filament[1]. Each tropomyosin molecule spans seven actin monomers within a filament and lays N- to C- terminally from actin's pointed to barbed end[2]. The 284 amino acid helix has a length of 420 Angstroms and has a molecular weight around 65-70 kilodaltons (vertebrate tropomyosin)[1][3]. A few of tropomyosin's characteristics as an actin binding protein includes regulation, stabilization and recruitment.
Tropomyosin's Structure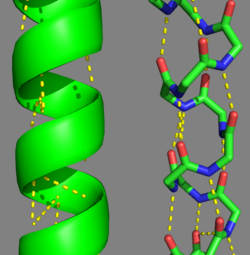 As determined by the Structural Classification of Protein's (SCOP) Database, Tropomyosin is categorized as follows (general to specific):
 Categorization of tropomyosin's describes a unique pattern of amino acids within the primary structure of the alpha helices that comprise the dimer interface[1]. The unique amino acid pattern, found within all coiled-coil proteins, is a heptad repeat, which follows a similar pattern to: H-P-P-H-C-P-C, where H is hydrophobic, P is polar and C is charged[1][3]. This heptad repeat forms a right-handed alpha helical secondary structure (see right for alpha helix secondary structure)[1]. This alpha helix is special in that it forms a hydrophobic strip along one side, which will interface with an adjacent alpha helix that also contains the heptad repeat and hydrophobic strip. These strips aid in the dimerization of tropomyosin and is important in the characteristic coiled-coil domain.
Post-Translational ModificationsThere are two types of post-translational modifications to tropomyosin: phosphorylation acetylation[1]. Phosphorylation occurs on amino acid [1]. This phosphorylation is occurs as a result of oxidative stress, which is associated with actin remodeling and recruitment of additional tropomyosin into stress fibers[1]. The acetylation occurs on the N-terminus of the N-terminal methionine, which is essential for: coiled-coil stability, overlap formation and actin binding[2][1]. Evolutionary ConservationTropomyosin is highly conserved actin binding protein, which is found in Eukarya from the animal kingdom to yeast, with the exception of plants[1]. The earliest characterization of tropomyosin gene lineage was in yeast (budding and fission yeast)[1][6]. These genes are known as TPM1 and TPM2, respectively, and share 64.5% sequence identity[1][6]. As we move away from unicellular organisms and into multicellular invertebrates, tropomyosin genes in nematodes slightly diverge from yeast, but have 85-90% sequence identity between their genes[1]. Further analysis for vertebrates show there are four genes that generate over 40 known mammalian isoforms of tropomyosin, which are synthesized by exon splicing[1]. The slight evolution change of tropomyosin has occurred as a result of the increasing need of tropomyosin to function in different systems, but tropomyosin has evolutionarily stayed well conserved because of the basic structural pressures imposed on the protein[1]. It is interesting to note, the region with the least conservation has been in the N and C terminus. This is the result of head-tail interactions changing to accommodate different polymer confirmations along actin for different functions[1]. (To see this evolutionary conservation, go to the top right image of the web page and click on the "show" link, which is to the right of "Evolutionary Conservation".) Tropomyosin's Function: Muscle and Non-Muscle SystemsMuscle tissue is comprised of many muscle fibers or cells. Those muscle fibers consist of myofibrils that contain a series of contractile units called sarcomeres. Within this unit contains thick filaments, comprised mainly of myosin, and thin filaments, which contain actin, tropomyosin and troponin. Tropomyosin in striated muscle systems (skeletal and cardiac) acts to inhibit the myosin cross-bridges from binding to the myosin binding site on thin filaments, this tropomyosin state is in the "Blocked" position[7]. When the muscle is stimulated, there is a rise in intracellular calcium stemming from a cascade of cellular processes. As the calcium is bathing the sarcomere, it will bind to the troponin complex, which is bound to both actin and tropomyosin[8]. The troponin will displace the tropomyosin from a "Blocked" to a "Closed" position[7]. This transition allows the myosin head to interact weakly with the myosin binding site[8]. The tropomyosin is displaced to its final position, "Open" state, along actin filament as myosin binds to its site[7]. These three tropomyosin states along the filament is referred to as the three state model[7]. As the intracellular calcium concentration falls, the troponin no longer is able to displace tropomyosin and it will transition back to the "Blocked" state[7].
3D structures of Tropomyosin
|
| ||||||||||
3D structures of Tropomyosin3D structures of Tropomyosin
Updated on 18-May-2022 {{#tree:id=OrganizedByTopic|openlevels=0|
- Tropomyosin
- 5kht – TPM α-1 N terminal - human
- 6otn - hTPM α-3 N-terminal
- 1c1g – pTPM – pig
- 6klq, 6klp - mTPM – mouse - Cryo EM
- 1mv4 - rTPM α-1 C-terminal - rat
- 2b9c – rTPM mid region
- 3azd – rTPM N terminal
- 3mtu, 3mud – cTPM α-1 – chicken
- 1ic2 - cTPM α-1 (mutant)
- 3u1a, 3u1c – cTPM α-1 N terminal
- 3u59 – cTPM β N terminal
- 7bjg, 7bjn – DmTPM 270-334 – Drosophila melanogaster
- 2tma – TPM - model
- 5kht – TPM α-1 N terminal - human
- Tropomyosin complexes
- 6kn8, 6kn7 - hTPM α-1 + troponin + actin – Cryo EM
- 6x5z - hTPM α-1 + myosin-7 + cActin – Cryo EM
- 5jlh - hTPM α-3 + myosin-14 + actin – Cryo EM
- 6g2t, 6cxj, 6cxi - hTPM + myosin-binding protein + actin – Cryo EM
- 7jh7 - pTPM + myosin-7 + actin – Cryo EM
- 7ko4, [[7kor5], 7ko7, 7kon, 7kor - pTPM α-1 + troponin + actin – Cryo EM
- 5nog, 5noj, 5nol - pTPM α-1 + actin – Cryo EM
- 2efr, 2efs, 2d3e - raTPM α-1 C-terminal+GNC4 leucine zipper – rabbit
- 4a7f, 4a7h, 4a7l - raTPM α-1 + myosin IE heavy chain + actin – Cryo EM
- 3j4k - raTPM + actin – Cryo EM
- 7nep - mTPM α-1 + myosin-4 + myosin light chain + actin – Cryo EM
- 7qin - mTPM α-1 in actomyosin complex – Cryo EM
- 3j8a, 5jlf - mTPM + actin - Cryo EM
- 2g9j - rTPM α-1 TM9A+GNC4
- 2w49, 2w4u – cTnnC+cTnnT+cTnnI+cTPM α-1 + cActin
- 7bjs – DmTPM 252-334 + kinesin heavy chain
- 2z5h – yTPM α-1 N-terminal+C-terminal+GNC4 leucine zipper+TnnT – yeast
- 2z5i - yTPM α-1 N-terminal+C-terminal+GNC4 leucine zipper
- 1kql - yTPM α-1 C-terminal+GNC4 leucine zipper
- 6kn8, 6kn7 - hTPM α-1 + troponin + actin – Cryo EM
}}
ReferencesReferences
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 1.18 1.19 1.20 1.21 1.22 1.23 1.24 Tropomyosins. I. Gunning, Peter, 1950- II. Series.[DNLM: 1. Tropomyosin. W1 AD559 v.644 2008 / WE 500 T856 2008]
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 Frye J, Klenchin VA, Rayment I. Structure of the tropomyosin overlap complex from chicken smooth muscle: insight into the diversity of N-terminal recognition . Biochemistry. 2010 Jun 15;49(23):4908-20. PMID:20465283 doi:10.1021/bi100349a
- ↑ 3.0 3.1 3.2 Whitby FG, Phillips GN Jr. Crystal structure of tropomyosin at 7 Angstroms resolution. Proteins. 2000 Jan 1;38(1):49-59. PMID:10651038
- ↑ 4.0 4.1 4.2 Clayton JE, Sammons MR, Stark BC, Hodges AR, Lord M. Differential regulation of unconventional fission yeast myosins via the actin track. Curr Biol. 2010 Aug 24;20(16):1423-31. Epub 2010 Aug 12. PMID:20705471 doi:10.1016/j.cub.2010.07.026
- ↑ 5.0 5.1 Stark BC, Sladewski TE, Pollard LW, Lord M. Tropomyosin and myosin-II cellular levels promote actomyosin ring assembly in fission yeast. Mol Biol Cell. 2010 Mar 15;21(6):989-1000. Epub 2010 Jan 28. PMID:20110347 doi:10.1091/mbc.E09-10-0852
- ↑ 6.0 6.1 Drees B, Brown C, Barrell BG, Bretscher A. Tropomyosin is essential in yeast, yet the TPM1 and TPM2 products perform distinct functions. J Cell Biol. 1995 Feb;128(3):383-92. PMID:7844152
- ↑ 7.0 7.1 7.2 7.3 7.4 Lehman W, Galinska-Rakoczy A, Hatch V, Tobacman LS, Craig R. Structural basis for the activation of muscle contraction by troponin and tropomyosin. J Mol Biol. 2009 May 15;388(4):673-81. Epub 2009 Mar 31. PMID:19341744 doi:10.1016/j.jmb.2009.03.060
- ↑ 8.0 8.1 Tyska MJ, Warshaw DM. The myosin power stroke. Cell Motil Cytoskeleton. 2002 Jan;51(1):1-15. PMID:11810692 doi:10.1002/cm.10014
