Byron's Bender: Difference between revisions
Eric Martz (talk | contribs) No edit summary |
Eric Martz (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
<table align="right" style="padding-left:10px;background-color:# | <table align="right" style="padding-left:10px;background-color:#e8e8e8;" width="420"><tr><td> | ||
[[Image:Byrons-bender-model.gif]] | [[Image:Byrons-bender-model.gif]] | ||
</td></tr><tr><td> | </td></tr><tr><td> | ||
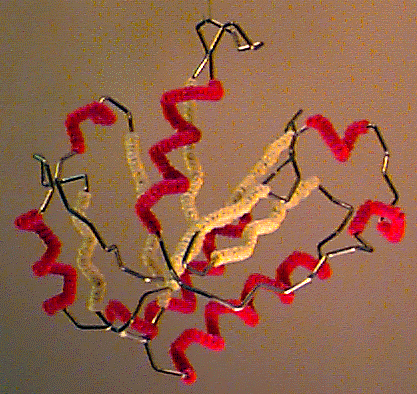
Byron's Bender wire model of the backbone trace of the I domain of CD11a (from [[1lfa]]), an alpha/beta domain of a leukocyte integrin adhesion molecule. Alpha helices are decorated with red pipe cleaners, while beta strands are decorated yellow. Model generously constructed by Tim Herman in 1997. | Byron's Bender wire model of the backbone trace of the I domain of CD11a (from [[1lfa]]), an alpha/beta domain of a leukocyte integrin adhesion molecule. Alpha helices are decorated with red pipe cleaners, while beta strands are decorated yellow. Model generously constructed by Tim Herman in 1997. | ||
</td></tr></table> | </td></tr></table> | ||
Byron's Bender<ref>Rubin, Byron; Richardson Jane S. The simple construction of protein alpha-carbon models. Biopolymers. 1972; 11(11):2381-5. [https://onlinelibrary.wiley.com/doi/epdf/10.1002/bip.1972.360111116 PDF]</ref><ref>Rubin, Byron. 1985. Macromolecule backbone models. ''Methods in Enzymology'' '''115''':391-7.</ref> is a machine designed to bend wire into the backbone trace of a protein model. Wire is advanced in fixed-length increments, representing the distances between alpha-carbon atoms. A dihedral angle is then set on dials, and a lever is pulled to bend the wire to the specified angle. Often, the emerging wire model collides with the machine. Then, the wire is cut, and later fastened end to end with a small metal sleeve. | Byron's Bender, invented by crystallographer Byron Rubin<ref>Rubin, Byron; Richardson Jane S. The simple construction of protein alpha-carbon models. Biopolymers. 1972; 11(11):2381-5. [https://onlinelibrary.wiley.com/doi/epdf/10.1002/bip.1972.360111116 PDF]</ref><ref>Rubin, Byron. 1985. Macromolecule backbone models. ''Methods in Enzymology'' '''115''':391-7.</ref>, is a machine designed to bend wire into the backbone trace of a protein model. The machine was manufactured by the Charles Supper Company in Natick Massachusetts<ref>[http://www.charles-supper.com/en/page/product.cfm?idProduct=33 Supper Protein Wire Model Bender], and widely used by crystallographers in the 1970's and 1980's. | ||
Wire is advanced in fixed-length increments, representing the distances between alpha-carbon atoms. A dihedral angle is then set on dials, and a lever is pulled to bend the wire to the specified angle. Often, the emerging wire model collides with the machine. Then, the wire is cut, and later fastened end to end with a small metal sleeve. | |||
==See Also== | ==See Also== | ||
Revision as of 00:50, 11 April 2018
|
Byron's Bender wire model of the backbone trace of the I domain of CD11a (from 1lfa), an alpha/beta domain of a leukocyte integrin adhesion molecule. Alpha helices are decorated with red pipe cleaners, while beta strands are decorated yellow. Model generously constructed by Tim Herman in 1997. |
Byron's Bender, invented by crystallographer Byron Rubin[1][2], is a machine designed to bend wire into the backbone trace of a protein model. The machine was manufactured by the Charles Supper Company in Natick Massachusetts<ref>Supper Protein Wire Model Bender, and widely used by crystallographers in the 1970's and 1980's.
Wire is advanced in fixed-length increments, representing the distances between alpha-carbon atoms. A dihedral angle is then set on dials, and a lever is pulled to bend the wire to the specified angle. Often, the emerging wire model collides with the machine. Then, the wire is cut, and later fastened end to end with a small metal sleeve.