1e2h: Difference between revisions
No edit summary |
No edit summary |
||
| (3 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
== | |||
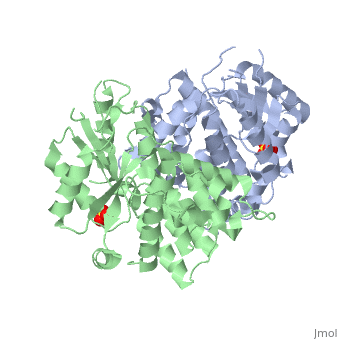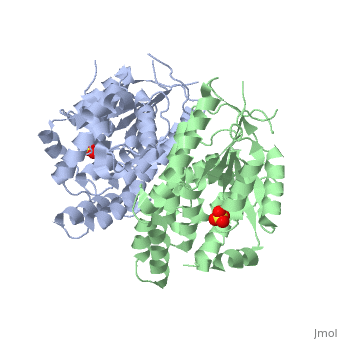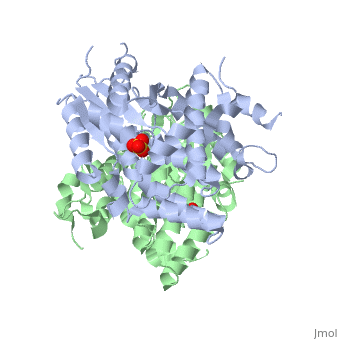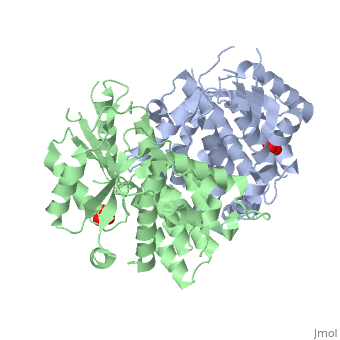
<StructureSection load='1e2h' size='340' side='right' caption='[[1e2h]], [[Resolution|resolution]] 1.90Å' scene=''> | ==The nucleoside binding site of Herpes simplex type 1 thymidine kinase analyzed by X-ray crystallography== | ||
<StructureSection load='1e2h' size='340' side='right'caption='[[1e2h]], [[Resolution|resolution]] 1.90Å' scene=''> | |||
== Structural highlights == | == Structural highlights == | ||
<table><tr><td colspan='2'>[[1e2h]] is a 2 chain structure with sequence from [ | <table><tr><td colspan='2'>[[1e2h]] is a 2 chain structure with sequence from [https://en.wikipedia.org/wiki/Human_alphaherpesvirus_1_strain_17 Human alphaherpesvirus 1 strain 17]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1E2H OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1E2H FirstGlance]. <br> | ||
</td></tr><tr id=' | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 1.9Å</td></tr> | ||
<tr id=' | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=SO4:SULFATE+ION'>SO4</scene></td></tr> | ||
<tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1e2h FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1e2h OCA], [https://pdbe.org/1e2h PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1e2h RCSB], [https://www.ebi.ac.uk/pdbsum/1e2h PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1e2h ProSAT]</span></td></tr> | |||
<tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | |||
</table> | </table> | ||
== Function == | == Function == | ||
[ | [https://www.uniprot.org/uniprot/KITH_HHV11 KITH_HHV11] In latent infection, may allow the virus to be reactivated and to grow in cells lacking a high concentration of phosphorylated nucleic acid precursors, such as nerve cells that do not replicate their genome (By similarity). | ||
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
Check<jmol> | Check<jmol> | ||
<jmolCheckbox> | <jmolCheckbox> | ||
<scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/e2/1e2h_consurf.spt"</scriptWhenChecked> | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/e2/1e2h_consurf.spt"</scriptWhenChecked> | ||
<scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
<text>to colour the structure by Evolutionary Conservation</text> | <text>to colour the structure by Evolutionary Conservation</text> | ||
| Line 31: | Line 31: | ||
==See Also== | ==See Also== | ||
*[[Thymidine kinase 3D structures|Thymidine kinase 3D structures]] | |||
*[[Thymidine kinase|Thymidine kinase]] | |||
== References == | == References == | ||
<references/> | <references/> | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
[[Category: | [[Category: Human alphaherpesvirus 1 strain 17]] | ||
[[Category: | [[Category: Large Structures]] | ||
[[Category: Scapozza | [[Category: Scapozza L]] | ||
[[Category: Schulz | [[Category: Schulz GE]] | ||
[[Category: Vogt | [[Category: Vogt J]] | ||
Latest revision as of 11:10, 6 December 2023
The nucleoside binding site of Herpes simplex type 1 thymidine kinase analyzed by X-ray crystallographyThe nucleoside binding site of Herpes simplex type 1 thymidine kinase analyzed by X-ray crystallography
Structural highlights
FunctionKITH_HHV11 In latent infection, may allow the virus to be reactivated and to grow in cells lacking a high concentration of phosphorylated nucleic acid precursors, such as nerve cells that do not replicate their genome (By similarity). Evolutionary Conservation Check, as determined by ConSurfDB. You may read the explanation of the method and the full data available from ConSurf. Publication Abstract from PubMedThe crystal structures of the full-length Herpes simplex virus type 1 thymidine kinase in its unligated form and in a complex with an adenine analogue have been determined at 1.9 A resolution. The unligated enzyme contains four water molecules in the thymidine pocket and reveals a small induced fit on substrate binding. The structure of the ligated enzyme shows for the first time a bound adenine analogue after numerous complexes with thymine and guanine analogues have been reported. The adenine analogue constitutes a new lead compound for enzyme-prodrug gene therapy. In addition, the structure of mutant Q125N modifying the binding site of the natural substrate thymidine in complex with this substrate has been established at 2.5 A resolution. It reveals that neither the binding mode of thymidine nor the polypeptide backbone conformation is altered, except that the two major hydrogen bonds to thymidine are replaced by a single water-mediated hydrogen bond, which improves the relative acceptance of the prodrugs aciclovir and ganciclovir compared with the natural substrate. Accordingly, the mutant structure represents a first step toward improving the virus-directed enzyme-prodrug gene therapy by enzyme engineering. Nucleoside binding site of herpes simplex type 1 thymidine kinase analyzed by X-ray crystallography.,Vogt J, Perozzo R, Pautsch A, Prota A, Schelling P, Pilger B, Folkers G, Scapozza L, Schulz GE Proteins. 2000 Dec 1;41(4):545-53. PMID:11056041[1] From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine. See AlsoReferences |
| ||||||||||||||||||